As we have seen over the past year, the ability to accurately predict the course of an epidemic is imperative when making decisions about how to control the spread of a virus. In this work, which focuses on influenza, Oxford Mathematicians Rahil Sachak-Patwa, Helen Byrne and Robin Thompson (now at the University of Warwick) investigated whether accounting for cross-immunity between related strains could improve forecast accuracy.
Influenza viruses mutate over time, and previous exposure to influenza confers partial protection against future infections with related strains. To understand whether including cross-immunity could improve forecasts of future epidemics, the team considered two ordinary differential equation models (Fig. 1).
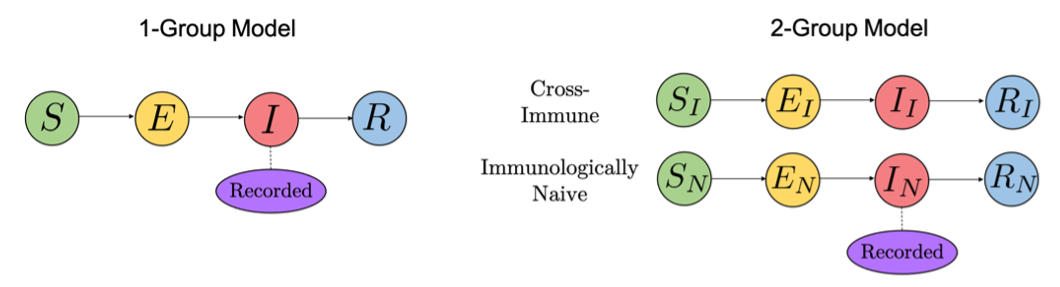
In both models, the population is partitioned into distinct classes which represent susceptible, exposed (i.e. infected but not yet infectious), infected, and recovered individuals. In the ‘1-group model’, all individuals are assumed to be identical and cross-immunity is neglected. In the ‘2-group model’, the population is split into two groups; cross-immune individuals who have previously been infected by a related strain, and immunologically naïve individuals who have no immunoprotection. It is assumed that the cross-immune individuals are less likely to experience severe disease and therefore recover more quickly than immunologically naive individuals, and hence only cases of naïve individuals get recorded.
Figure 1 – Schematic of the two models.
The team fitted both models to estimated case notification data from the 2009 H1N1 influenza pandemic in Japan, and then generated synthetic data for a future outbreak by assuming that the 2-group model more accurately represents the influenza infection dynamics. They then used both models to generate real-time forecasts for future outbreaks as they occur. Model parameter values estimated from the 2009 epidemic were used as informative priors, motivated by the fact that without using prior information from 2009, the forecasts are highly uncertain.
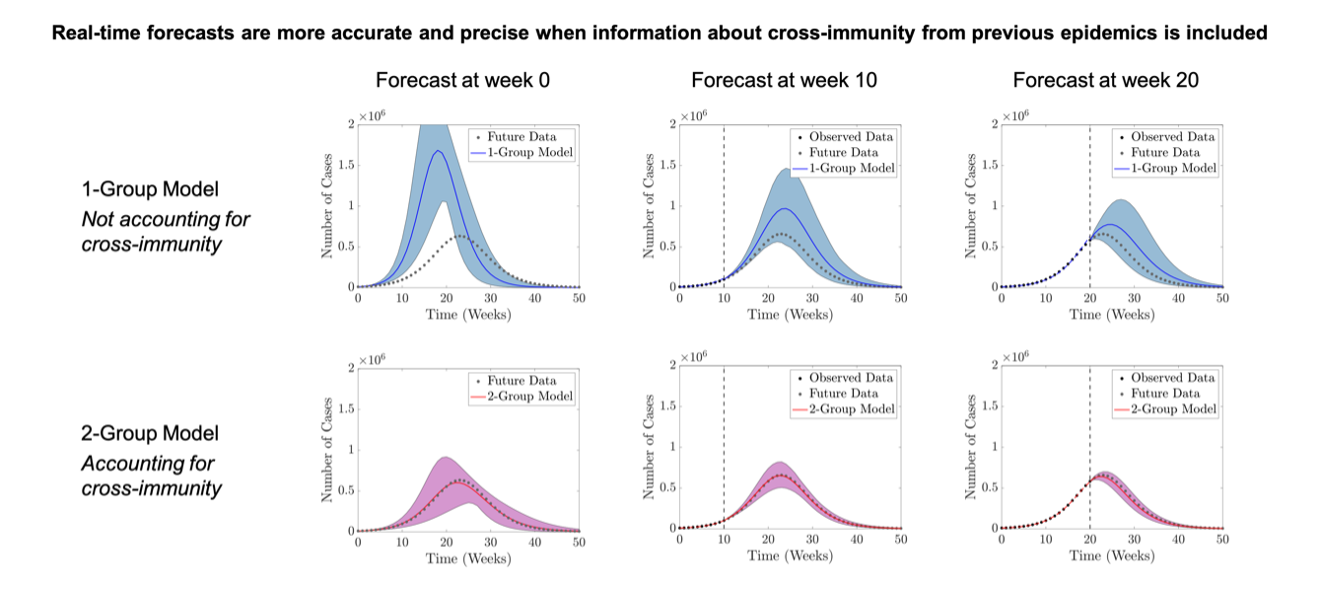
In the scenario that was considered for an epidemic occurring 25 years into the future, the 1-group model only produced accurate outbreak forecasts once the peak of the epidemic had passed, while the 2-group model made accurate forecasts early on (Fig. 2). Based on these results, the team conclude that accounting for cross-immunity, driven by exposure to previous epidemics, improves the accuracy of forecasts during influenza outbreaks. These findings could also be applied to the ongoing coronavirus pandemic, where incorporating information about cross-immunity between strains into models could result in better predictions about the spread of the virus.
Figure 2 – Forecasts of a future epidemic occurring 25 years into the future using the 1-group and 2-group models.