West Nile virus (WNV) is responsible for viral encephalitis in humans, a condition that causes inflammation of the brain and can have longer-lasting physical effects. WNV is also related to similar viruses such as Dengue and Zika that are also of significant public health concern. Faced with the challenge of understanding how the virus reproduces within the host and its potential for epidemic, Oxford Mathematician Soumya Banerjee and colleagues have developed a computational method to determine characteristics of WNV infection even in the face of limited experimental data, an approach that could be applicable to other emerging diseases like the Zika virus for which there is little data.
Diseases that jump the species barrier from animals to humans affect millions worldwide. An understanding of how disease progression and immune response vary from species to species could have important public health benefits. In their work the team attempt to understand how immune function scales with body size, work which is a foundation for understanding scaling of immune response to other pathogens or in other animals.
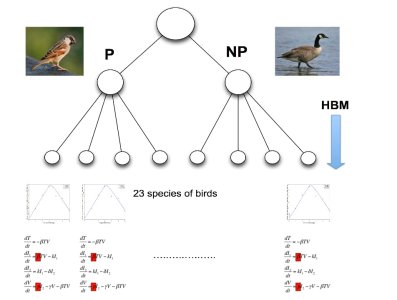
The team’s computational framework for infectious disease modelling at the within-host level leverages data from multiple species. This is likely to be of interest to modellers of infectious diseases that jump species barriers and infect multiple species – the method can be used to determine computationally the competency of a host to infect mosquitoes that will sustain West Nile virus infection. The models show that smaller Passerine species are more competent in spreading the disease than larger non-Passerine species. This suggests the role of host phylogeny as an important determinant of within-host pathogen replication.
Ultimately the team believes their work could be an important step in linking within-host viral dynamics models to the between-host models that predict spread of infectious disease between different hosts.