How does cellular metabolism change in different environments? Metabolism is the result of a highly enmeshed set of biochemical reactions, naturally amenable to graph-based analyses. Yet there are multiple ways to construct a graph representation from a metabolic model.
This work, a collaboration between Oxford Mathematician Mariano Beguerisse and colleagues in mathematics and bioengineering from Imperial College and the Polytecnic University of Valencia, proposes a principled framework to construct genome-scale models of metabolism using modern network science. These models resolve various challenges, such as the incorporation of pool metabolites, directionality of metabolic flows, and scenario-specific flux information.
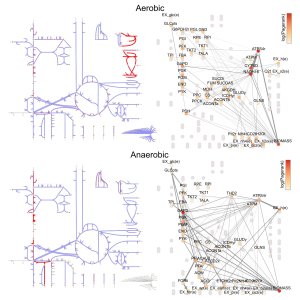
This framework abandons metabolic descriptions that are generic blueprints in favour of tailored metabolic descriptions under any specific context of interest. For example, this model predicts the way in which the metabolism of Escherichia coli re-routes metabolic flows as environmental conditions change from being oxygen-rich (aerobic) to oxygen-poor (anaerobic). In many situations the reactions that constitute metabolic pathways form tighly-knit clusters, but in other cases the reactions may be dispersed, or not even connected to the network. Creating scenario-specific metabolic networks allows us to study how pathways behave in different conditions, and understand the role that individual reactions play. An analysis of the metabolism of human liver cells with a rare metabolic disease identifies several key reactions that flux-only analyses miss because they do not incorporate the rewiring of the metabolic network.
The method can be integrated into pipelines based on flux balance analysis and provides a systematic framework to explore changes in network connectivity as a result of environmental shifts or genetic perturbations. The paper giving more details on the research appears in the journal NPJ Systems Biology and Applications.
--
The image above shows two metabolic reaction networks of E. coli. The top network shows the metabolic connectivity under normal conditions. When the oxygen is removed from the environment, the cell must drastically re-wire its metabolism in order to survive (bottom). Click to enlarge.